Labeled dataset of X-ray protein ligand images in 3D point cloud and validated deep learning models
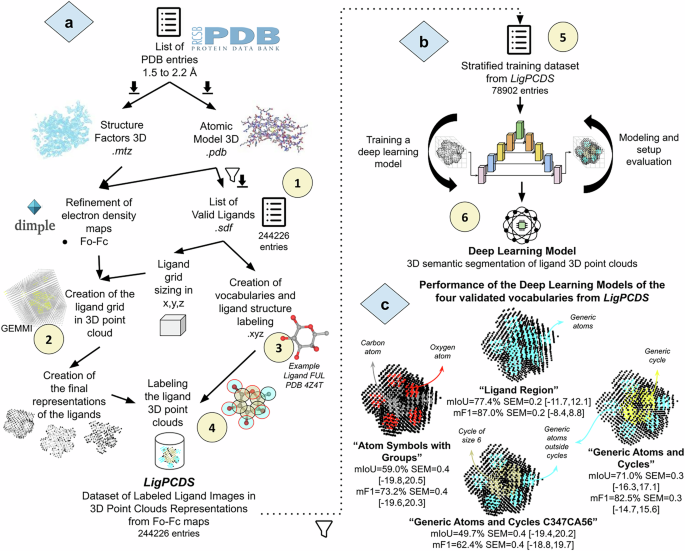
The LigPCDS dataset creation followed six major steps (Fig. 1), which are summarized below and explained in detail in the next subsections.
-
1.
Creation of a list of valid ligands from RCSB PDB.
-
2.
Creation of the representations of the ligand 3D image in 3D point clouds.
-
3.
Creation of chemical vocabularies and ligand structure labeling.
-
4.
Labeling ligand 3D point clouds.
-
5.
Creation of a stratified training dataset from LigPCDS.
-
6.
Training, optimization and validation of DL models.
The validation steps (steps 5 and 6 in Fig. 1b,c) of LigPCDS methodology are presented in the Technical Validation section.
Hardware
The hardware used to execute the LigPCDS creation and the DL models training is a computer with the following configuration: AMD Ryzen 9 3950X CPU, 16 cores and 32 threads, 128 Gb RAM and 2x GeForce RTX 2080 SUPER GPUs with 8 Gb of dedicated RAM each (hardware A). Exceptions were for specific DL models analyses that are point out in the text and used hardware B, a cluster with the following configuration: AMD EPYC 7742 CPU with 64 cores and 80 threads, 384 Gb RAM and 4 GPUs NVIDIA HGX A100 with 40 Gb of dedicated RAM each.
List of valid ligands
To obtain a list of ligands (step 1, Fig. 1), the advanced search tool of the RCSB PDB ( was initially used to retrieve all entries with resolution between 1.5 Å and 2.2 Å, in December 2019. The chosen resolution range aligns with the most frequent resolution values found in the PDB (Supplementary Figure 1) and those typically obtained in structural biology and drug discovery projects. Additional selections to the retrieved RCSB PDB files were: the presence of free ligands (non-covalent), availability of experimental data (entries with electron density maps also deposited), data originated from X-ray experiments with proteins, and deposited at PDB after January 2008 (more stringent validation metrics in PDB). For the free ligands, we have selected organic molecules formed by atoms of carbon, oxygen, nitrogen, phosphor, sulfur, iodine, fluorine, chlorine, bromine or selenium; hydrogen atoms were omitted here due to their poor detection by X-ray crystallography at the chosen resolution range. At this stage, this resolution range would reduce data variations caused by large differences in resolution for LigPCDS construction, while keeping ligand information that is still difficult to predict. Other ranges were not tested so far, and may be used in the future.
A total of 39,353 PDB entries were selected using the above criteria, containing 13,189 unique ligand codes (unique ligand structure). The.pdb and.mtz files of these RCSB PDB entries were downloaded automatically. The coordinate lines representing the ligands present in the protein chains of these PDB entries were isolated from the retrieved files and saved into individual.pdb files. This procedure resulted in a total of 293,822 available ligand entries from 39,169 PDB entries, containing 13,074 unique ligand codes.
The Structure Data Format (SDF) file of each ligand entry was also downloaded from RCSB PDB. An SDF file is a chemical file format for molecular data based on the MOL-file format – which can store single or multiple molecules, describing all their atoms in 3D coordinates. Each ligand’s SDF file was used to build and validate the ligand representative molecular graph (chemical structures). The free ligand entries with validated SDF files were used to propose chemical vocabularies for labeling the structure of protein ligands using a building block-like approach. This structure validation resulted in a total of 259,606 ligand entries from 39,052 PDB entries, containing 12,972 unique ligand codes.
To validate the experimental data of each PDB entry, a standardized procedure was proposed to refine the datasets downloaded from RCSB PDB (.mtz and.pdb files), without the ligand atomic entries, aiming to improve the blob imaging and to remove any failed PDB entry (described in the next subsection). In addition, the ligand entries with validated SDF files were also used to extract the ligand’s 3D representations from their correctly refined Fo-Fc maps (described in the next subsections). The ligand entries that raised an error in any step were removed from the list of valid ligands.
The final list of valid ligands contains 244,226 entries of ligands from 36,202 PDB deposits. These ligands represent non-covalent protein ligands composed by C, O, N, P, S, Se, F, Cl, Br and/or I atoms, where 12,239 are unique ligand codes (unique structures) with frequencies ranging from 1 to 33,063 occurrences (20 ± 526). Single atoms or ions (e.g. Cl-) correspond to 8.6% of the ligand entries (n = 21,003), while the other 91.4% are valid molecular structures (n = 223,223). The median size of valid ligands is 6 atoms and the mean size is 11 non-hydrogen atoms, with sizes ranging from 1 to 140 non-hydrogen atoms. These statistics indicate a great imbalance problem in the list of valid ligands, which is related to the diversity of non-covalent ligands deposited in PDB. They also highlight the diversity of potential protein ligands with importance in biology and drug discovery. Many of such ligands are still to be discovered and will have to be interpreted in the future, as novel X-ray protein structures in complex with ligands are obtained.
The RCSB PDB downloads were automated with Python v3.8 scripts, and the ligand entries validation used the functionalities of the RDKit package v2019.09.3 ( 16.9% of the ligand entries and 8% of the PDB entries were excluded during validation, 11.6% of the ligand entries due to invalid SDF files (minor download errors are also included), 4.0% due to refinement errors and 1.3% due to errors in the creation and labeling of the ligand’s 3D representation. This indicates poor quality of part of the ligand entries, further highlighting the difficulties for directly applying data mining techniques on PDB data19.
Ligand 3D representation in point cloud
Next in LigPCDS creation, the 3D representations of the ligands present in the list of valid ligands were designed and created. Considering the variability and flexibility in the size and conformation of ligands, the ease and speed of manipulating point clouds29, and the availability of many good performance deep learning architectures for 3D point clouds30, we have chosen point clouds as the format to represent the 3D images of ligands in LigPCDS.
The point clouds were initially extracted from the Fo-Fc maps using a ligand grid. For this, a 3D grid box was drawn around the ligand and the electron density intensity values in each x,y,z coordinate of the grid was computed and stored in the color channels of the point cloud. Then, contours and scales were applied to extract the 3D representations of the ligand images, without background and noise. Nine types of 3D representations (at different contours and scales) were generated to each ligand and are available at LigPCDS. The representation type to be used in a given application will depend on the desired application of the user, in a case-by-case basis. For our deep learning model of ligand chemical structure prediction, the qRankMask_5 representation showed the best results.
The detailed schema used in LigPCDS for creating the 3D representations of ligands in 3D point cloud format (step 2, Fig. 1) is shown in Fig. 2. A step-by-step explanation of this process is given below.
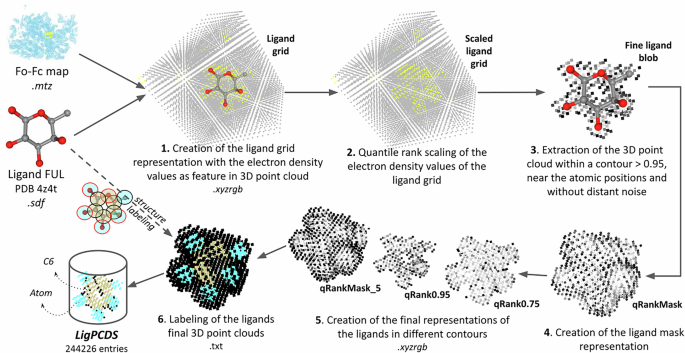
Schema for creating the labeled representations of ligands in 3D point cloud format for LigPCDS. The ligand FUL of PDB (entry 4Z4T) was used to exemplify the creation of the ligand’s 3D point cloud starting from the grid up to the final 3D representations. (1) The ligand’s grid representation is sized and interpolated from its Fo-Fc map in all its x,y,z positions, using the Gemmi package26. The ligand’s grid is stored in point cloud format (.xyzrgb) with the density value of each point saved in its RGB channels (feature as colors). (2) The density values of the ligand’s grid 3D point cloud are transformed and normalized using the quantile rank scale33. (3) The points of the ligand’s grid within a contour of 0.95 (value > 0.95) are selected and only the points near the ligand’s atomic positions and closely connected (with a distance between points smaller than grid space * 1.42 + 0.15) are retained, the rest is removed as noise. This creates the fine ligand blob representation. (4) The ligand’s mask point cloud is created from this fine ligand blob by applying a 1.1 Å radius expansion from its borders and is named “qRankMask”. (5) The final representations of the ligands are created by applying different contours in the ligand’s mask representation and extracting the selected 3D point cloud. The final representations are named as “qRank” followed by the contour value, e.g. “qRank0.95”. Additionally, a representation equal to the ligand’s mask and with all values below 0.5 set to zero is created and named “qRankMask_5”. This schema corresponds to the procedures used to complete step 2 in the LigPCDS creation workflow (Fig. 1a). (6) Finally, the labels of the ligand’s structure are used for pointwise labeling the final 3D representations of the ligands, which corresponds to step 4 of the LigPCDS workflow (Fig. 1a).
Refinement of the Fo-Fc maps (experimental data preparation)
Before extracting the 3D representations of the ligand’s blob in 3D point clouds, each PDB entry in the list of valid ligands were first refined using the Dimple software v2.6.1 ( a macromolecular crystallographic pipeline for refinement incorporated into the CCP4 program suite25. A standardized Dimple refinement was performed for each PDB entry using their respective downloaded.mtz and.pdb files, with the option of removing heteroatoms (it removes all ligands from the.pdb file) and with two refinement cycles (longer refinement). The other parameters of Dimple received their default values. Dimple refinement was carried out with two primary objectives: first, to highlight the presence of any ligand blob in the crystal structure. With the “remove heteroatom” parameter active, the unmodeled electron density related to the ligands (high values in the Fo-Fc maps) could be revealed, and any bias related to incorrect ligand structure modelling on the PDB deposit would be removed. Second, to improve the overall Fo-Fc map and the local quality of the ligand blob, further normalizing the model refinement standards for the different crystal structures present in the list of valid ligands. The PDB entries that presented errors in the refinement were excluded. The list of valid ligands at this point contained 36,325 PDB entries successfully refined, with 247,878 ligand entries listed, from which 12,250 were unique ligands.
Extraction of the ligand grid representation in 3D point cloud (procedure 1, Fig. 2)
A ligand grid was then created to extract the 3D image of each ligand blob (found in the refined Fo-Fc map) into the 3D point cloud format. The ligand grid is a bounding box defined on the boundary of the ligand’s atomic positions, plus a gap, designed to cover the complete shape of the ligand blob. This procedure used the original SDF coordinates of the ligand to locate the center of its molecular structure in the refined Fo-Fc map, and to retrieve the ligand’s atomic 3D coordinates, thus computing the bounding box on the boundary of its atomic positions. Through experimental inspection, this box was expanded with an additional gap equal to 4.2 Å in its boundaries (equal to the diameter of the largest theoretical radius31 – Supplementary Table 1), and then, a second 120% expansion of its size was performed. The obtained dimensions defined the size of the ligand grid in the Fo-Fc map, centered on the ligand boundary box.
The Gemmi package26 v0.5.8 was then used to interpolate the values of the Fo-Fc map for all x,y,z positions of the ligand grid. The obtained 3D grid was stored in a point cloud format, named the ligand grid representation. The difference electron density value of each point was chosen as the feature for the ligand 3D representation. The interpolated density value of each point (feature) was stored in the color channels of the 3D point clouds of the ligand grid representation. A spacing equal to 0.5 Å for the points of the ligand grid was tested and chosen. This value is smaller than the distance of a chemical bond (a sigma C-C bond measures around 1.54 Å) and allows to retain more details in the final 3D representations.
The Gemmi v0.5.8 Python package26 for structural biology provides a framework of functions to manipulate electron density maps in indexable 3D grids, behaving like standard numerical vectors. Gemmi v0.5.8 allows extracting 3D grids from specific regions of an electron density map with different spacing between the points. It uses an implementation of the trilinear interpolation of the 8 closest points32 of a given position of a map to compute its electron density value.
Transformation and scale of the ligand grid representation (procedure 2, Fig. 2)
The quantile rank scale was then used to transform and scale the ligand grid to allow for their correct comparison. This is an equivalent approach to histogram equalization33,34 in image processing. This scale normalizes the values in the range from 0 to 1. The quantile rank scale is used in other crystallography applications33, and replaces the density value ρ(x,y,z) of each point by its position in the quantile distribution of the points for the region being considered. This scale does not change the shape of the electron density, all points that have the same ρ density values have the same value in this function. Furthermore, unlike the sigma scale, which must be applied globally across the entire electron density map, the quantile rank scale can be applied locally within a box to compare the same region. The sigma and quantile rank scales are comparable, with 1σ, 2σ or 3σ contours corresponding to quantile positions that vary approximately between 0.85, 0.95 and 0.9833. The use of the quantile rank scale allows to speed up calculations for data extraction, improves comparison, and excludes noise from the electron density map of distant regions, since the resolution of X-ray protein crystallographic data varies locally35.
A fast implementation of the quantile rank scale function was created for this project: first it sorts the density values inside the ligand grid representation and then replaces the value of each point by its position in the ranked quantile distribution of the 3D-grid. Ties receive the first occurring position to the left. The scaled ligand grid representation for 247,424 ligand entries, 12,245 being unique ligands, were successfully created at this step.
Extraction of the fine ligand blob 3D representation (procedure 3, Fig. 2)
The next step consisted in removing noise from the scaled ligand grid. For this, the scaled ligand grid representation was filtered to retrieve only the points within a contour of 0.95 (value > 0.95). Then, only the points near the ligand atomic positions and closely connected (with a distance between points smaller than the grid space × 1.42 + 0.15) were retained. By applying a neighborhood searching approach it was possible to remove the noisy points filtered from the ligand grid representation at 0.95 contour; in other words, the points that were not closely connected to the ligand atomic positions were removed here. This created the fine ligand blob 3D representation with a strong signal level and without noise. Python’s Open3D package29 v0.12 functionality was used to create the 3D point cloud of the ligand grid, mask and final representations (described in the next section). This package has an implementation of KDTrees using the FLANN library36 for quick access of the closest neighborhood of the point clouds. This allowed searching with good performance.
Creation of the ligand mask representation (procedure 4, Fig. 2)
The fine ligand blob 3D representation at 0.95 contour was then used as a reference for the blob location and shape. This 3D representation was expanded from its boundary points with a radius equal to 1.1 Å in the scaled ligand grid. The resulting 3D point cloud was stored as the final ligand mask representation and was named qRankMask. By doing this expansion on the “fine ligand blob 3D representation”, instead directly on the scaled ligand grid representation at 0.95 contour (no filters), we could prevent distant noisy points from being included in the qRankMask and further in the final representations of the ligands.
Creation of the final representations of the ligands in 3D point cloud (procedure 5, Fig. 2)
Finally, the 3D representations of the list of valid ligands in 3D point cloud were created. Nine types of 3D representations were generated per ligand entry by exploring different contour levels. All of them compose LigPCDS. The representation types were named: qRank0.5, qRank0.7, qRank0.75, qRank0.8, qRank0.85, qRank0.9, qRank0.95, qRankMask, and qRankMask_5. These fine sliced 3D point clouds were obtained by applying, to the ligand mask representation (qRankMask), contours at 0.5, 0.7, 0.75, 0.8, 0.85, 0.9 and 0.95 on the quantile rank scale. The different contours used are related to the representation name suffix. These point clouds have as a single feature the scaled density value of the qRankMask normalized again from 0 to 1, where each contour value is the new 0 in the final representation. For qRankMask_5 a different approach was used, aiming to join types qRank0.5 and qRankMask which gave better results in the models training: values below 0.5 were set to 0 in the qRankMask, and all the normalized values of contour 0.5 were directly used as feature. In other words, week points (below 0.5) were clipped.
The ligand mask representations (qRankMask and qRankMask_5) and the representations with a quantile rank contour ≤ 0.8 (qRank0.5, qRank0.7, qRank0.75, qRank0.8) gave better results when training the validated deep learning models, with a very small difference between their accuracies. The representation qRankMask_5 was chosen as the best result for the validated segmentation models; it maintains the ligand mask shape with good accuracy. Depending on the usage goals of this dataset, different representation types may give the best results.
A total of 244,283 ligand entries, 12,239 being unique ligands, had their final 3D representations successfully created. The first and fourth columns of Fig. 3 show the final 3D point clouds of two different ligands in four different representation types and the ligand mask. This figure illustrates the impact of the contour value on the final 3D point cloud of the ligands.
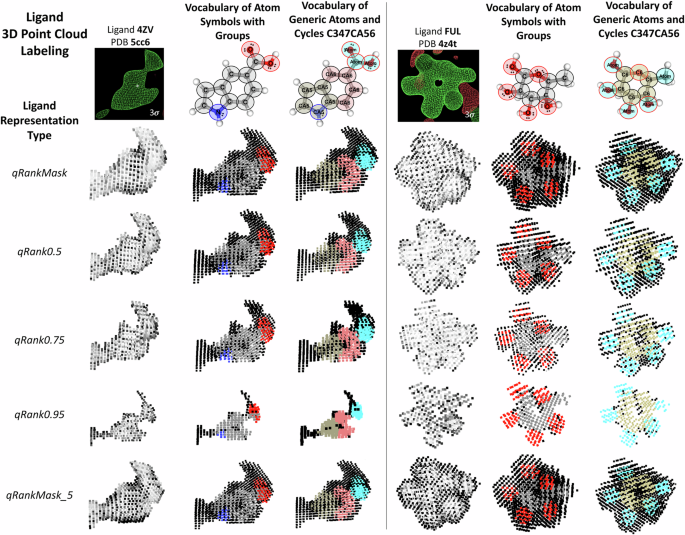
Example of a ligand’s 3D point cloud labeling for five different representation types. Two ligands are used for illustration: 4ZV (PDB entry 5cc6, resolution 2.1 Å) and FUL (PDB entry 4z4t, resolution 1.8 Å). Their blobs from their Fo-Fc maps are shown in the top of the panel with a contour of 3σ (image created with Coot). The LigPCDS visualization script was used to draw the ligands’ 3D point clouds. For ligand FUL, it is possible to see the pattern of a ring in the qRank0.95 representation; it results from the cyclic substructure of size six, present in its structure. In ligand 4ZV this pattern is not clear, possibly due to the mobility of this molecule – which is indicated by the presence of noise around its image (blob) and its representations (bottom left and top right of the ligand region – black points labeled as background). Furthermore, the qRank0.95 representation of ligand 4ZV is partially fragmented, with missing points, while for ligand FUL all points with labels are completely covered. There is more visual correspondence between the ligand’s image in the 3σ Fo-Fc maps and the qRank0.95 point cloud.
The mean time to create the ligand grid representation in 3D point cloud was 0.33 seconds per ligand. The mean time to create all representation types was 0.39 seconds per ligand (mean time for a spacing of the points equal to 0.5 Å). Other ways to create the 3D representations of ligands in 3D point clouds may also be tested in the future. This work provides one of the possible frameworks of functions to create 3D representations of protein ligands in 3D point clouds (imaging approach), which were successfully tested to be used in ML approaches.
Chemical vocabularies and ligand structure labeling
Chemical vocabularies were designed (step 3, Fig. 1) to compose the building blocks to label the created 3D representations of ligands in 3D point clouds from LigPCDS. The set of uniquely used labels is referred to as vocabulary and the unique labels are referred to as classes.
Data labeling can be very difficult depending on the amount of data and on the availability of validated references37. The labeling in LigPCDS was designed to first label the ligand’s structure atom-wise with building blocks (classes) and then to extrapolate it to the ligand 3D representations (the ligand chemical structure – next subsection). The implemented structure labeling approach was inspired by ML solutions that model chemical structures of small molecules for drug design38.
Four simplified chemical vocabularies were designed and validated (please see Technical Validation section) for labeling the ligand’s structure (Table 1). They are based on the atom’s symbol (the atom itself), which represent the individual scattering contribution of each atom to the electron density map; and on cyclic structures information, which adds a layer of 3D spatial distribution and geometrical restrains for the ligand region, and consequently to the blob region. All vocabularies also contain the background class, which represents non-atom regions of the ligands, and is only used in the labeling of the ligand 3D point cloud.
The four valid vocabularies designed are simplifications of two major labeling approaches: i) the AtomSymbol-based, with the chemical symbol of organic atoms (e.g. C, O, N, P, S, Se, Br, Cl, F, I); and ii) the SP-based, with the SP hybridization attributed to each atom (e.g., sp, sp2, sp3, sp3d1, sp3d2, sp3d3), which is defined by the atom steric number. The cyclic structure arrangement information is also included in both Atom Symbol and SP hybridization labeling. Please refer to Supplementary Note 1 for more information about the process in designing the chemical vocabularies. A brief explanation of the four valid vocabularies, which are directly mapped from the major labeling approaches, is given below and is summarized in Table 1:
-
I)
“Vocabulary of the Ligand Region” (SP-based, 2 classes): labels all atoms with the generic atom class;
-
II)
“Vocabulary of Generic Atoms and Cycles” (SP-based, 3 classes): labels the atoms as generic atoms outside cyclic structures and atoms in generic cyclic structures (of any size and type);
-
III)
“Vocabulary of Generic Atoms and Cycles C347CA56” (SP-based, 9 classes): labels the atoms as generic atoms outside cyclic structures and atoms in cyclic structures with sizes (ranging from 3 to 7), where cyclic structures with sizes 5 and 6 are further labeled according to their aromaticity (aromatic or not). Aromatic cyclic structures of sizes 4 and 7 are not distinguished from non-aromatic ones due to their low abundance. Cyclic structures with more than 7 atoms are not distinguished from atoms outside cyclic structures as large cyclic arrangements are more flexible and may not have a shape pattern in the Fo-Fc map;
-
IV)
“Vocabulary of Atom Symbols with Groups” (AtomSymbol-based, 6 classes): labels the ligand atoms with their chemical symbol, if it is one of the most common atom symbols in organic molecules (C, O, N); or with the following groupings: the “halo” group, if it is a halogen atom (atom symbols F, Cl, Br and I), and the “PSe” group, if it is one of the remaining atoms with lower abundance in the dataset (atom symbols P, S and Se).
The ligand structure labeling procedure was automated in a Python script with the RDKit package v2019.09.3 and was used to implement both the AtomSymbol-based and SP-based approaches. It works as follows. For each ligand: (i) all cyclic structures in the ligand structure are retrieved; (ii) for each atom of the ligand, its label is set to its SP hybridization (one of sp, sp2, sp3, sp3d, sp3d2, sp3d3), or its atom symbol (one of C, O, N, P, S, I, F, Se, Cl and Br), depending on the parameters. This label is concatenated with the smaller cyclic structure in size and aromatic cyclic arrangement type in which this atom appears (one of C3, CA4, C4, CA5, C5, CA6, C6, CA7 or C7 in this order), if any. Finally, (iii) the labels of all atoms are returned. The labels are mapped to the atoms using their unique coordinates in the 3D space.
These two major approaches (AtomSymbol-based and SP-based) were used to label the structures of the ligands in the list of valid ligands, resulting in 244,226 ligand structures successfully labeled. The ligands structural labeling results were saved to tables in .xyz files (CSV format), with one atom per row and their information and label by column. These results were stored in the xyz directory of the data record of each major approach: SP-based and AtomSymbol-based labeling (detailed in the Data Records section). The mapping from the two major approaches to the four validated vocabularies was performed by matching their labels with the provided mapping tables presented in Supplementary Tables 2, 3 (see Usage Note for more details). Examples of structure labeling with these two major approaches and their four mapped and validated vocabularies are illustrated for the molecules beta-L-fucose and 1H-indole-5-carboxylic acid, which have the following ligand codes in PDB: FUL and 4ZV, respectively (Fig. 4).
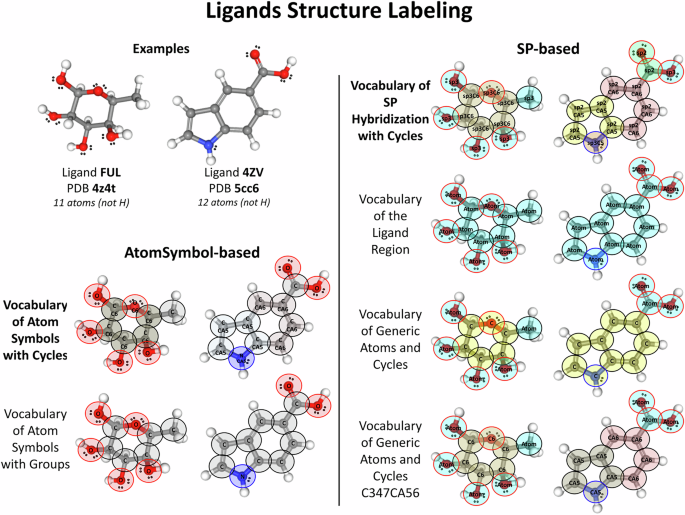
Examples of a ligand’s structure labeling. Ligands 4ZV and FUL from PDB (entries 5CC6 and 4Z4T, respectively) are shown on the top left panel and were used as example to illustrate all the proposed vocabularies: the “Vocabulary of SP hybridization with Cycles” and its three mappings (SP-based approach), which are shown on the right panel; and the “Vocabulary of Atom Symbols with Cycles” and its mappings (AtomSymbol-based approach), which are shown on the bottom left panel. The label of each atom is written inside its atomic sphere (represented by a circle), which is colored according to its label in the filling and the border color received the atom color in the 2D structure.
The four valid vocabularies are further described with the distribution of occurrences of their classes by atom in the final list of valid ligands (Figs. 5 and 6). These distributions help visualize the class imbalance problem39,40 present in LigPCDS, a crucial information to understand its limits for semantic segmentation tasks. Also, the maximum imbalance ratio40 (dmax, Eq. 1) was computed to indicate, for each vocabulary, the maximum level imbalance across classes and to help the comparison of the viability of the different vocabularies.39,40
$${d}_{\max }=\frac{\mathop{\max }\limits_{i}\left\{{C}_{i}\right\}}{\mathop{\min }\limits_{i}\left\{{C}_{i}\right\}}$$
(1)
where \({C}_{i}\) is the number of atoms labeled as class \(i\), and \(\mathop{\max }\limits_{i}\left\{{C}_{i}\right\}\) and \(\mathop{\min }\limits_{i}\left\{{C}_{i}\right\}\) are the maximum and minimum number of labeled atoms among classes, respectively.
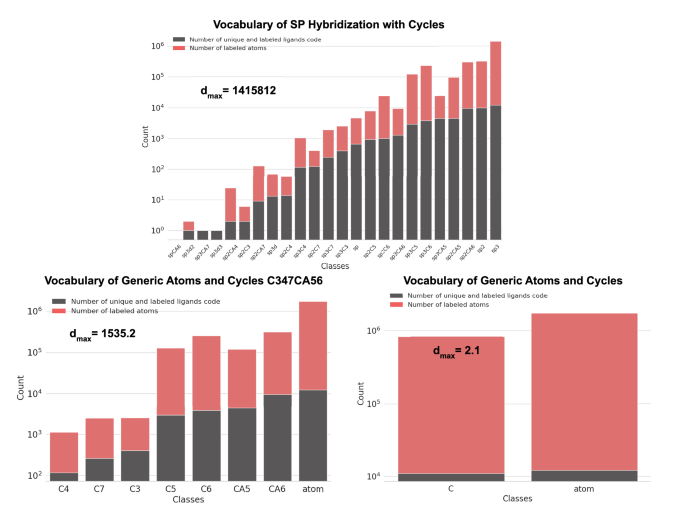
Class distribution of SP-based vocabularies. Distribution of the class occurrence in the SP-based vocabularies by labeled atom of all entries of the final list of valid ligands. Their corresponding imbalance ratio (dmax) is also presented. The distribution for the “Vocabulary of the Ligand Region” is omitted because all 2,566,614 atoms were labeled with the same generic class of atoms. The background class is not used in these distributions.
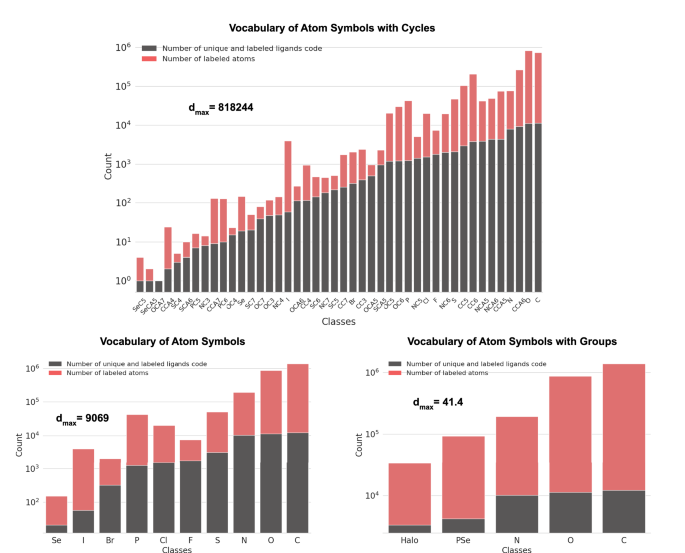
Class distribution of AtomSymbol-based vocabularies. Distribution of class occurrence in the AtomSymbol-based vocabularies by labeled atom of all entries of the final list of valid ligands. Their corresponding imbalance ratio (dmax) is also presented.
Figure 5 displays the distribution of class occurrences using the SP-based vocabularies for the atoms of the ligands in the list of valid ligands. Figure 6 displays this distribution for the AtomSymbol-based vocabularies.
The two vocabularies that kept more chemical information and had good accuracy in relevant classes of the validated models (please see Technical Validation) were selected as the best labeling approaches: the “Vocabulary of Generic Atoms and Cycles C347CA56” and the “Vocabulary of Atom Symbols with Groups”.
Labeling of the final representations of the ligands in 3D point clouds
The last step to obtain LigPCDS (step 4, Fig. 1; procedure 6, Fig. 2) is the pointwise labeling of the final representations of the list of valid ligands in 3D point clouds. This was done with the atom-wise extrapolation of the labels of the ligands’ structures (previous section) to their final representations in 3D point clouds.
A widely used model to calculate the atomic volume of molecules is to treat atoms as rigid spheres41. These spheres have a radius equal to the van der Waals theoretical atomic radius for each atom type, and serve as a model to represent the electron density volume that would be occupied by each atom of the molecule. The electron density is theoretically distributed as a Gaussian centered on each atom41, with high intensity values at the center. When a contour is applied to the electron density (e.g. in the sigma or quantile rank scale), only the central peak of each Gaussian is visible42. Batsanov’s work31 summarizes the available data on the van der Waals theoretical atomic radius for molecules and crystals. The work that describes XGen42, for fitting ligands in the real space of electron density maps, provided information for the typical experimental X-ray radius for organic elements at different experimental electron density resolutions.
It was thus decided to use the modeling of an atomic sphere to extrapolate the labeling from the atoms of the ligands structure to their final 3D point clouds, using as radius 65% of the experimental radius provided by XGen for each atom type. This percentage was chosen to recover the central region of the density peak of each atom, while keeping the contour of the ligand’s structure. The resolution of the PDB entries was used to select the sets of radii for each ligand entry, rounding the resolution to the first decimal place (values tabled for resolutions 1.5, 1.6, 1.7, 1.8, 1.9, 2.0, 2.1 and 2.2 Å in Supplementary Table 1). For Selenium (Se) atom, which does not appear in the XGen table, it was assigned the radii of the Bromine (Br) atom. The points in the representation of the ligands that are not covered by the atomic spheres with 65% of the experimental radius of XGen received the labeling of background noise (“background” class – regions in the Fo-Fc map without a ligand atom). Points in the intersection region of two or more atomic spheres received the label of the nearest atom center. Other percentages of atomic radii were not tested. This procedure was implemented with the functionality of the Open3D v0.12 library for quick access of the neighborhood of each point.
The ligand’s structure labeling was extrapolated to the final 3D representations of the ligands present in the list of valid ligands using the two major labeling approaches (SP-based and AtomSymbol-based). A dataset of labeled 3D representations of the difference electron density of ligands in point cloud was obtained for each major vocabulary. The ligand final 3D point clouds that were correctly labeled and tested constitute 244,226 entries in the final list of valid ligands. The point cloud labeling testing is detailed in the Technical Validation section.
The labeled records of ligand images in 3D point cloud representations were called “LigPCDS-SP” and “LigPCDS-AtomSymbol”, which correspond to the SP-based and AtomSymbol-based labeling approaches, respectively, and compose LigPCDS. This dataset covers entries of free protein ligands of organic molecules (non-covalent protein ligands composed by C, O, N, P, S, Se, F, Cl, Br or I atoms), obtained from X-ray protein crystallography, with experimental resolutions ranging from 1.5 to 2.2 Å. These records (SP-based and AtomSymbol-based) are organized by PDB entry and contain all the final 3D point clouds of the list of valid ligands that appear in the respective entry. The organization of this dataset is detailed in the Data Records section. Two examples of final labeled 3D point clouds of ligands with the “Vocabulary of Generic Atoms and Cycles C347CA56” and the “Vocabulary of Atoms Symbols with Groups” are presented in Fig. 3 for different representation types.
link





.jpg)