A chemoenzymatic reaction-diffusion system with local inhibition and distal activation
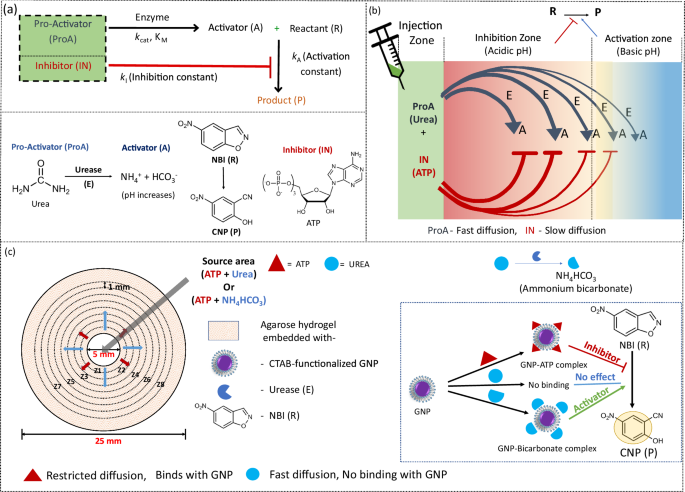
Designing of the experimental framework
Our chemoenzymatic RD system is based on a catalytic hydrogel matrix with a reaction-diffusion system (Fig. 1). As mentioned in the preceding section, we aimed in activating a reaction spatially distant from the region of addition of inputs. We chose a base-catalyzed proton-transfer ring-opening reaction (also known as Kemp Elimination (KE) reaction) for this purpose56,57,58,59. The substrate for KE reaction 5-nitrobenzisoxazole (NBI, denoted as R in the manuscript) was evenly distributed in the agarose gel matrix. This R is converted to 2-cyanonitrophenol (CNP, denoted as P) in presence of a base and the reaction gets catalyzed by cationic gold nanoparticle (GNP)56,57. Therefore, the hydrogel was also embedded with GNP. Notably, in ultrapure water (with a resistivity of 18.2 MΩ·cm at 25 °C), GNP does not catalyze the conversion of R to P. An enzyme, urease (E), is also present in the gel matrix that does not react with R or GNP. E can convert a neutral molecule, urea (dormant activator or pro-activator, ProA) to a base, ammonium bicarbonate (activator, A). We named urea and ammonium bicarbonate as ProA and A, respectively as A can activate the conversion of R to P and A formed only after the enzymatic action over ProA37. As we aimed to develop a LIDA-based system, we chose an anionic molecule, ATP as inhibitor (IN) of the KE reaction. We locally injected a mixture of ProA (urea) and IN (ATP) to achieve our desired system with local or short-range inhibition and distal or long-range activation.
The reasons behind choosing ATP as IN and urea as ProA are following – (i) due to acidic nature of ATP, it is known to suppress KE reaction, and thereby, it acts as a very good inhibitor for this process. Secondly, for local inhibition, IN needs to be slow to diffuse. In our case, ATP is anionic and has a great ability to bind with cationic GNP (already embedded in gel matrix), therefore, due to binding, diffusion of IN will be restricted through the gel matrix. (ii) We chose urea as ProA. As it is neutral, it does not bind to GNP and diffuses faster than IN. Therefore, upon injecting a mixture of ProA and IN, enzymatic ProA to A conversion will take place even in a zone devoid of IN. Thus, in principle, R to P conversion will take place far away from the zone of injection of ProA and IN. However, there is always the question of whether adding A (instead of ProA) and IN would provide the same effect or not. To address this issue, we also performed control experiments both experimentally and theoretically to demonstrate that enzymatic conversion of ProA to A during diffusion is a pre-requisite for developing such a system in a well-defined manner.
For experimental verification of the above-mentioned hypothesis, we designed our system as shown in Figs. 1c and 2a. In a petri dish, agarose gel casting was performed by keeping the center with 5 mm diameter vacant. The height of the gel was 3 mm. In the vacant zone, a solution of ATP + urea or ATP + NH4HCO3 was added and allowed to diffuse through the gel matrix. The gel is also embedded with cationic CTAB-capped gold nanoparticles (GNP), urease (E) and NBI (R). We used 0.75 wt% agarose gel for all our experiments for the following reasons (Supplementary Fig. 1). We found at higher concentration of agarose gel (1 wt %) GNP gets aggregated which can decrease the NBI to CNP reaction rate and alter the diffusion behavior of ATP. The aggregation was checked by noting the ratio of absorbances A620/A520, where an increase in the ratio indicates more aggregated AuNPs60. Higher agarose concentrations create a gel with a smaller mesh size, which acts as a physical barrier to nanoparticle movement, resulting enhanced interparticle friction and clustering of nanoparticles61. In contrary, at lower agarose concentration (0.5 wt%), the time taken to complete gel formation (after dissolving agarose at 110 °C) was ~25–30 min, resulting in some background conversion of NBI to CNP, prior to the addition of any reagent at the center. Therefore, to balance both issues, we used 0.75 wt% agarose gel for our study, where GNP aggregation was negligible and gel formation was fast (~5 min). We would like to mention here that only after complete solid-like gel formation, the solution of ATP + urea or ATP + NH4HCO3 at the center was added. Additionally, while preparing gel, we did not use any salt solution, as salt like NaCl can result in GNP aggregation and inhibit the base-catalyzed NBI to CNP reaction (Supplementary Fig. 2).
Interplay of urea and ATP to govern final Product (P) formation kinetics within the chemoenzymatic reaction systems in water
Before proceeding to the experiment with gel matrix, we analyzed the system in aqueous solution (milli Q water). At first, we checked the binding of ATP and NH4HCO3 and non-binding of urea with cationic GNP surface via zeta potential measurement (Supplementary Fig. 3). Then we checked in the presence of urease, addition of urea indeed results in conversion of NBI to CNP and the time of reaction outcome depends on the amount of urea (Supplementary Fig. 4). In absence of urease, no reaction occurs, and reaction can get activated in presence of NH4HCO3 (Supplementary Fig. 5). It is worth mentioning that in our previous studies, we already demonstrated the catalytic ability of GNP for this KE reaction in urea-urease system33. We also showed that with increasing ATP content in the reaction system with urea-urease and NBI, the KE reaction rate can be significantly slowed down (Supplementary Fig. 4). For example, only 0.2 mM ATP can slow down the NBI to CNP conversion rate by 50% when 2 mM urea was used in presence of 15 nM urease. In a similar condition, upon checking the inhibitory effect of a simple salt NaCl, we found 10–15 mM NaCl is needed to decrease the production of CNP by 50% (Supplementary Fig. 2). In this case, weakly basic chloride ion can act as a competitor of basic HCO3– ion for binding with the cationic GNP surface to hinder the conversion of NBI to CNP. Using a pH probe, bromothymol blue (BTB), we established that with increasing ATP, the time required for acidic to basic transformation was increased, resulting in a delay of the base-catalyzed proton transfer reaction (Supplementary Fig. 6)56,62.
Spatiotemporal evolution of the final product due to the RD process in the gel matrix
Now, for the gel experiment, we measured the formation of CNP at 8 different zones, each separated by 1 mm in a plate reader (please see experimental procedure in the methods section for details). First, we injected 40 μl volume of aqueous solution containing 50 mM urea and 5 mM ATP at the center and allowed it to diffuse through the hydrogel matrix (Figs. 1c and 2a). We then monitored the evolution of reaction product, P by following the absorbance at 380 nm wavelength over time at different points (Fig. 2b, c). Upon injecting 50 mM urea + 5 mM ATP at the center, we were pleased to observe that the formation of P did not happen near the injection center (Fig. 2d, Supplementary Fig. 7). Indeed, in this case, the maximum product was found 5 mm away from the center between 2 h to 5 h (Fig. 2e). For the initial 60 and 90 min data, we observed that the maximum product formation was taking place at zone 4 (4 mm away from center). Interestingly, while injecting 40 μl of 50 mM NH4HCO3 (product formed after urea-urease reaction) + 5 mM ATP at the center, the maximum reaction product formed in the very beginning at zone 3 (up to 60 min), and then the maximum product always formed at zone 2 (Fig. 2f, Supplementary Fig. 8). Although in this case, the reaction rate was very high and the amount of product was almost equally distributed from zone 1 to zone 5, 6 and 8, after 1, 2 and 4 h, respectively, suggesting the difference in concentration is negligible and the product mostly formed near the injection center. We also performed an additional experiment with lower urea (25 mM) but with similar ATP of 5 mM. In this case, we found the maximum product forming zone was further away by 1 mm up to 90 min in comparison to 50 mM urea + 5 mM ATP case (Fig. 2e, Supplementary Fig. 9). At both 1 and 1.5 h, the maximum product formation was found at zone 5 which was maintained up to 5 h. Again, at fixed ATP (2.5 mM), and decreasing urea from 25 to 10 mM, the maximum product formation zone moved further away at zone 5, compared to zone 4 after 3 h (Fig. 2f, Supplementary Figs. 10, 11). These experiments clearly suggest the importance of injecting urea, instead of NH4HCO3 and enzymatic conversion of urea to NH4HCO3 during diffusion plays the key role in generating distant reaction product forming zone. Also decreasing the amount of urea injected at a fixed ATP concentration, resulted in the product formation zone moving further away from reaction center. However, at very low urea (10 mM), the amount of formed P was too low. We also performed control experiments without either urease or GNP or both in the gel matrix. In all these cases, the product formation was too low, indicating the requirement of both E and GNP for the reaction conversion (Supplementary Figs. 12–14). Next, we also checked the effect of addition of only urea in the system having GNP and urease in the gel matrix (Supplementary Fig. 15). Here also, the product formed maximally at zone 1 and diffused to other distant zones with time, showing the importance of ATP (IN) and conversion of urea (ProA) to basic ammonium ions (IN) during diffusion through gel matrix for LIDA-effect.
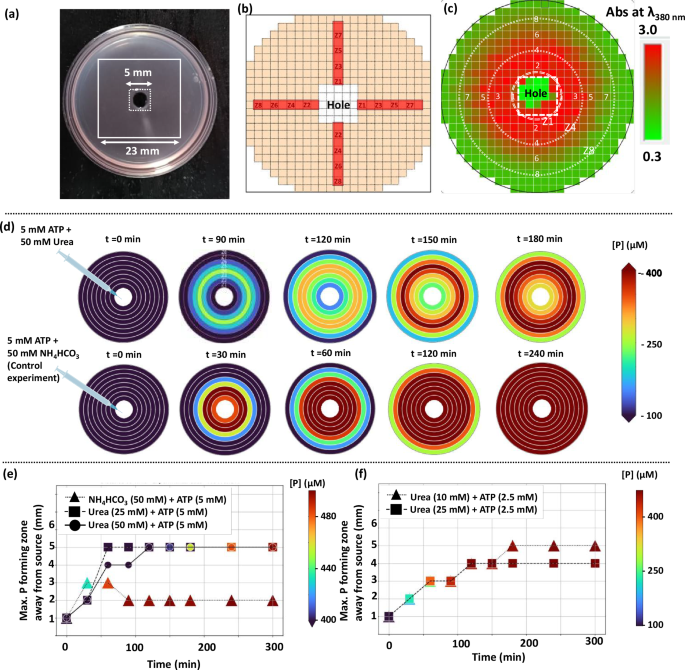
a Snapshot of the experimental petridish with dimension of the experimentally measured area (square with solid line) and hole with dashed square. b Schematic representation of the points taken to average out the value of each zone (shown by the red highlighted parts). c A snapshot of the experimental data inside plate reader to show the hole and other experimental zone. Measurements were taken in all directions. To avoid the error due to angular contribution, the average values of the four points (as shown in cross pattern in panel b) at the same distance were calculated and extrapolated to represent the entire circumference as circle (indicated by showing dotted circle to represent zone 1 (Z1), zone 4 (Z4) and zone 8 (Z8) in panel c). d Spatiotemporal pattern of product (CNP) formation when 40 µl of aqueous solution of 5 mM ATP and 50 mM urea (top panel) and 5 mM ATP + 50 mM NH4HCO3 (bottom panel) were injected at the central zone. The color map showed the CNP concentration at each zone. Herein, concentrations above 400 µM were shown as deep red and below 100 µM as deep blue. e The maximum product-forming zone away from the source at different time points was plotted for the injection of urea (25 mM) + ATP (5 mM), urea (50 mM) + ATP (5 mM), and NH4HCO3 (50 mM) + ATP (5 mM). The color of each symbol depicts the concentration range as shown in the color map (below 400 µM as deep blue and 500 µM as deep red). f Maximum product forming zone away from the source at different time points was plotted for injection of urea (25 mM) + ATP (2.5 mM) and urea (10 mM) + ATP (2.5 mM). The color of each symbol depicts the concentration range as shown in the color map. Data are presented as mean ± SD (where n = 8). Source data are provided as a source data file.
Spatiotemporal change in pH of the LIDA-based chemoenzymatic systems
The results discussed in the previous paragraph suggest that enzymatic conversion of ProA to A during diffusion is the essential factor for developing LIDA-based systems. Next, we were curious to find the reason behind this phenomenon. As KE is a base-catalyzed reaction, R to P conversion should occur at high pH. Therefore, we were interested to find out the spatiotemporal evolution of pH in the system after injection of 50 mM urea and 5 mM ATP. For this, bromothymol blue (BTB) dye was incorporated in the gel instead of NBI, along with GNP and E (Fig. 3a). At first, we calibrated the gel system at 620 nm wavelength with BTB dye using known pH value (Supplementary Fig. 16). It is worth mentioning that absorbance value at 620 nm increases with increasing pH. Interestingly, we find that the initial zones (up to 2 mm) near the injection center remain acidic (below pH 6) for about 60 min, whereas pH of zone 3 and 5, at 60 min were 7.2 and 8.1, respectively (Fig. 3b, c, Supplementary Figs. 17, 18). It clearly suggests that initially the pH remains low at the injection center but was higher at zone 4 and 5. Even after 180 min, we found pH of zone 1 was close to 7, whereas the value was 8 or higher for zone 4 onwards. As pH of the distant zone becomes higher quickly, the reaction starts at distal zones from the injection center (Fig. 3d, Supplementary Fig. 17). This also suggests that acidic ATP remains close to injection center, whereas urea moved faster and converted to basic NH4HCO3, even at distant zones to activate the KE reaction.
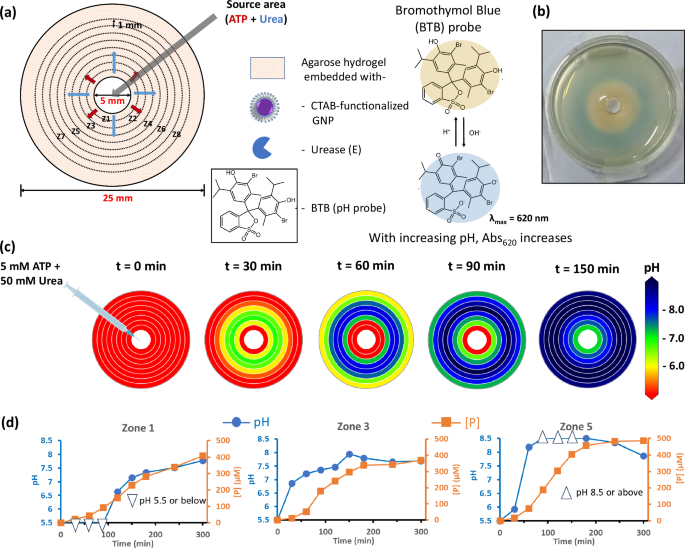
a Schematic of the reaction set up with components as described in Fig. 1c. In this case, pH probe Bromothymol blue (0.02 mM) was added in the gel instead of NBI. At the empty circular zone (5 mm diameter) aqueous solution of (ATP + urea) was added and allowed to diffuse to the agarose gel matrix (0.75 wt%). b A snapshot of petridish taken after 60 min, showing the initial zone is acidic (yellow color) and outer region is basic (blue color). c Spatiotemporal pattern of pH in the gel matrix with time when 40 µl of aqueous solution of 5 mM ATP and 50 mM urea was injected in the central zone. Blue indicates pH of 8 or higher and red indicates pH lower than 6. The pH of the whole gel before injection was around 5.6 as it was made with mQ water (which is slightly acidic due to dissolved CO2). d pH (blue points, left sided Y-axis) and product formation (orange points, right-sided Y-axis) with time at zone 1, zone 3 and zone 5. Data are presented as mean ± SD (where n = 8). Source data are provided as a source data file.
Numerical simulation of the LIDA-based model systems with different reaction and diffusion parameters
After establishing the system experimentally, we were interested to explore the spatiotemporal emergence of the reaction product at different diffusion parameters, activation and inhibition constant by developing a model system. This will be of importance for further understanding the present system and how to build similar LIDA-type synthetic system, with chemical components of different diffusivity and catalytic properties. This model simulates a two-dimensional reaction-diffusion system involving six chemical species undergoing transport and/or enzymatic reactions. The system is defined in polar coordinates \((r,\,\theta )\) over a circular domain, and it accounts for both radial and angular diffusion (Fig. 4). E and R are embedded in a circular reaction matrix. ProA and IN are injected at the center and allowed to diffuse and react. We also performed control experiments with only A instead of ProA to see the difference in spatiotemporal reaction pattern.

a Scheme of the reactions used for modeling. Here similar dimensions of two-dimensional circular disc (8 mm diameter) as used in experiment were chosen. b Spatiotemporal distribution of reaction product (P) at different kIN/kA and different diffusion coefficient ratios of ProA, IN and A when ProA (50 mM) and IN (5 mM) are injected at the center (left) and A (50 mM) and IN (5 mM) are injected at the center (right). The maximum product concentration (normalized) obtained in both ProA + IN and A + IN cases are given in Supplementary Table 1. Product distribution when kIN/kA = 103, and DProA😀IN😀A = 40: 1: 1, 40: 2: 2, 40: 5: 5 and 40: 8: 8 (where DProA = 4 × 10−10 m2/s) for c ProA + IN injection and d A + IN injection at the center at 150 min. Notably, product concentration in the case of A + IN injection is higher than ProA + IN injection. For (b–d), the concentration of product was normalized with respect to the maximum product concentration obtained for the case of kIN/kA = 103 and DProA😀IN😀A = 40:2:2 at 150 min.
The governing reaction-diffusion is described by partial differential equations (PDE) of the form:
$$\frac{\partial {C}_{i}}{\partial x}={D}_{i}{\nabla }^{2}{C}_{i}+{R}_{i}(C)$$
(1)
Where \(i\) is the appropriate chemical species ProA, A, I or P. \({C}_{i}\) is the concentration of the chemical species \(i\). \({D}_{i}\) is the diffusion coefficient. \({R}_{i}(C)\) represents the reaction term. A constant concentration of enzyme (E) in the matrix is assumed.
$${\nabla }^{2}{{\rm{is}}}\; {{\rm{the}}}\; {{\rm{Laplacian}}}\; {{\rm{in}}}\; {{\rm{polar}}}\; {{\rm{coordinates}}}:{\nabla }^{2}C=\frac{1}{r}\frac{\partial }{\partial r}\left(r\frac{\partial C}{\partial r}\right)+\frac{1}{{r}^{2}}\frac{{\partial }^{2}C}{\partial {\theta }^{2}}$$
(2)
The reaction terms for: ProA → A (Michaelis-Menten equation) and R → P conversions are given in Eqs. (3) and (4), respectively.
$${R}_{A}=\frac{{k}_{{cat}}{C}_{{ProA}}{C}_{E}}{{K}_{M}+{C}_{{ProA}}},{R}_{{ProA}}=-{R}_{A}$$
(3)
$${R}_{P}={{k}_{A}{C}_{A}R}_{0}\exp \left(-{k}_{A}{C}_{A}-{k}_{{IN}}{C}_{I}\right)t$$
(4)
Where \({R}_{0}\) is the initial concentration of the reactant, NBI.
In Eq. (4), kA and kIN are rate constant of A-induced activation and IN-mediated inhibition (see supplementary information for detailed derivation of Eq. (4)). Central Finite Difference Approximations are used for spatial derivatives. Neumann boundary condition (no-flux) is imposed at the center and domain boundary. A circular disc of 8 mm diameter has been chosen to follow the reaction-diffusion phenomenon, and we followed the RD system until 150 min. At first, we checked the spatiotemporal emergence of the final product P in different ratio of kIN and kA (kIN / kA = 1, 103 and 106) while adding a mixture of 50 mM ProA and 5 mM IN at the center of the circular disc. Additionally, we checked 4 different diffusion ratio of ProA, A and IN (DProA: DIN: DA = 40: 1: 1; 40: 2: 2; 40: 5: 5 and 40: 8: 8) (Fig. 4, Supplementary Figs. 19–30). Here, we assumed DProA = 4 × 10−10 m2/s (considering diffusion co-efficient of urea) and kcat/KM = 3000 mM−1s−1 (kcat = 104 s−1, KM = 3.3 mM) for enzyme urease towards substrate urea63. For screening of the simulation parameters we assume both IN and A has similar diffusion co-efficient as both can bind with GNP. The diameter of GNP is around 25 nm; therefore, we assumed that bound IN and A would be of similar diffusion co-efficient with a much lower value than ProA. Additionally, we did not consider any diffusivity of P. Simultaneously, we checked the scenario when instead of ProA, we added 50 mM A (directly added without allowing it to form through enzymatic reaction) and 5 mM IN with similar kinetic and diffusion parameters (Fig. 4, Supplementary Figs. 31–42). From the theoretical plot, we can observe a trade-off between diffusion ratio and kIN/kA. In general, for kIN/kA of 103 a clear LIDA effect was observed at all ratios of diffusion coefficients studied in this work. However, the amount of P steadily decreased with increasing diffusion of IN. After 150 min, the highest LIDA was observed in the case of kIN/kA of 106, with DProA: DIN: DA = 40: 2: 2, although the product amount was lower than the case when kIN/kA was 103. Similarly, the system will be effective for DProA: DIN: DA = 40: 5: 5 or 40: 8: 8 cases quite efficiently when kIN/kA equals 103, but not with kIN/kA of 106. In fact, we were pleased to observe while injecting only A = 50 mM and IN = 5 mM at the center, and DIN = DA (1:1), the product always formed close to the injection center and the amount of product was higher than the case when (ProA + IN) was injected (Fig. 4b–d, Supplementary Table 1).
After analyzing the above data, we fixed DProA: DIN: DA = 40: 2: 2 and kIN/kA to 103 to explore the spatiotemporal emergent behavior of P with different concentration ratios of ProA and IN injection at the center. This model output is comparable to our observed experimental result (Supplementary Figs. 43, 44). We fixed the amount of IN to 5 mM and decreased the amount of ProA to 25 and 10 mM. In this case we found that, with low ProA (10 mM), maximum P emerged at a distant zone (3 mm away from the center) beforehand (in terms of time) than in the other two cases (Supplementary Fig. 45). In a similar manner we fixed the amount of ProA to 50 mM and found that, with increasing IN, maximum P was formed at a distant zone at earlier time point compared to low IN (Supplementary Fig. 46). Lower ProA or higher IN results in enhanced inhibition, therefore, maximum product was found at more distant zone. However, with high IN or low ProA, the amount of product is low and in some cases, negligible. Both simulations are similar to our experimentally observed scenario described at Fig. 2f to a considerable extent. Again, while using only (A + IN) injection, the maximum product forming zone remains same irrespective of the concentration and no concentration effect over spatiotemporal variation was observed (Supplementary Fig. 47).
At this point, we asked ourselves, in the development of LIDA type system, what is the benefit of including ProA to A conversion and higher diffusion co-efficient of ProA compared to IN? In fact, one might think that this system could be developed using a faster diffusing A and slower diffusing IN. We tested this by simulating the A + IN system, assuming DA to be greater than DIN. The model simulated conditions when DIN😀A = 1:40, 2:40, 5:40 and 8:40 at kIN/kA = 1, 103 and 106, with the amount of A and IN being 50 mM and 5 mM respectively (fixed DA = 4 × 10−10 m2/s) (Supplementary Figs. 48–59). The resulting plot of P concentration over time shows that P formation was concentrated in the central zone, near the injection point at all diffusion coefficient ratios at kIN/kA = 1. At kIN/kA = 103, maximum concentration of P was found at zone 3 at all ratios of DIN😀A. P formation in the case of kIN/kA = 106 was maximum at distal zones (zone 5, 6), although the maximum concentration of P was lowest here, out of the three kIN/kA cases tested (Supplementary Fig. 60). With faster diffusion of IN, the zone where maximum P is observed moves further away from the site of injection. And higher kIN/kA value shows stronger inhibition in the central zones. Although LIDA can be observed in this case, the addition of ProA to A reaction gives greater spatiotemporal control over the system, even after considering ProA converted A has similar diffusivity to IN. To prove this, we compared these results with the earlier case of ProA + IN at various ratios of diffusion coefficients and kIN/kA (Supplementary Fig. 61). While the concentration of P is lower in the case of ProA + IN, the zone of maximum P formation is consistently farther away than the case with A + IN at least by 1 mm in each case. This is applicable at all ratios of diffusion coefficients and kIN/kA. We also compared P output pattern for two different situations, DProA: DIN: DA = 40:2:5 and 40:2:20 with DIN: DA = 2:5 and 2:20 cases (Supplementary Fig. 62). Here we found that addition of ProA results shifting of maximum P formation zone by 1–2 mm distance compared to A + IN case with different spatiotemporal manner. Overall, the results show the LIDA effect is even more effective while using ProA and its conversion to A during diffusion, even when A has higher diffusivity than IN.
One other advantage of having ProA is that temporal control can be achieved by modifying the reaction parameters of ProA to A. We simulated cases where we modified the kcat/kM ratio to increase or decrease by 10- and 100-fold (at a range of five orders of magnitudes from 30 to 300000 mM−1s−1) (Supplementary Figs. 63–68). The point of time when the desired concentration of P is observed in the system is tracked. The results show that temporal control of this RD system can be achieved by modifying reaction kinetics of ProA to A reaction. This temporal control would not be possible in a simple system with only two diffusing species as shown in A + IN case and therefore, validating the necessity of reaction-diffusion for more distinct LIDA with manifold spatiotemporal diversity.
Numerical simulation considering pH sensitivity of urease reactivity
So far, we have not considered the pH sensitivity of urease reactivity. Urease activity followed a bell-shaped curve where at pH 7 enzyme showed its highest activity. In our experiment, we found that the initial zones remain acidic due to presence of ATP, whereas outer zone has higher pH (data demonstrated in Fig. 3). Indeed, upon measuring ammonia production rate during urease-urea reaction, we also found that with increasing ATP concentration urease reactivity decreased (Supplementary Table S3). Here pH of the system is also decreasing with increasing ATP.
For modeling pH-dependent urease activity, a Gaussian curve centered at pH 7 (enzyme’s optimal pH) was used as previously shown by Straube et al.64. The experimentally measured pH values at each of the 8 zones over time are associated with the equivalent radial zones of the model at the specified time points (Supplementary Table 3). Zone 0, representing the central cavity is assigned a pH value of 3.7 to represent its acidic nature due to presence of 5 mM ATP. For each zone, the pH data is interpolated over the experimental time range using the ‘interp1d’ function of Python, so that the local pH can be looked up at any simulation moment for each radial position.
The pH in each zone determines the enzyme activity factor (activity_factor variable in code ProA+I – with pH dependence.py), a scalar between 0 and 1. This value then multiplies the catalytic reaction term (reaction_AB variable in the code) in the model equations, thus controlling how efficiently the substrate is converted into the product based on the local pH. Therefore, this model accurately captures the spatial and temporal progression of the reaction system, considering the effect of pH on urease activity. In this case we also considered diffusion of P which is similar to IN and A (DProA: DIN: DA: DP = 40: 2: 2: 2). The simulation result is given in Supplementary Fig. 69. In this case also a clear LIDA effect was observed for ProA + IN case compared to A + IN injected scenario with a factor of 1–2 mm.
In summary, firstly, we experimentally demonstrated a complex chemoenzymatic system to gain control over catalysis at a spatially distant zone in a time-dependent manner. This reaction-diffusion system primarily relied on supramolecular interaction-based diffusion control and both enzymatic and nanoparticle surface-mediated catalysis. This demonstration will be important in the future development of chemical systems that aim to regulate the onset of chemical reactions both spatially and temporally65,66,67,68,69,70. Notably, our approach is entirely different for modulating chemical reactions at the far-off site from the injection point. We also designed a theoretical model to rationalize the experimental facts. We further analyzed the benefits of injecting (dormant activator + inhibitor) together rather than (activator + inhibitor) both in terms of gaining efficient spatial gating of the reaction activation zone from the injection center and temporal control via tuning the enzymatic rate of conversion of activator from dormant activator during diffusion. Overall, pH enhancement and corresponding reaction activation region in the gel matrix can be spatiotemporally gated by the enzymatic conversion rate of ProA to A and initial source concentration of ProA and IN. We believe both experimental and theoretical data presented in this manuscript will be important for the advancement of synthetic reaction-diffusion systems to achieve spatiotemporally diverse chemical signaling pathways via modulating chemical inputs, their interaction parameters and directions of reaction propagation ability with surrounding environmental precursors (for example in presence of ions)71,72,73. One can also envision the utility of this approach from the triggering of spatially distant self-assembly and positional information based chemical events akin to artificial developmental program for the construction of more complex chemical neural networks47,74.
link





.jpg)