Every researcher knows the frustration: your groundbreaking experiment is on hold because the DNA sequence you need is “unmanufacturable”. Or worse, after weeks of waiting, the oligos arrive with errors that send you back to square one.
These aren’t isolated incidents, they are symptoms of an outdated DNA synthesis paradigm that’s increasingly misaligned with modern research demands. Enzymatic DNA synthesis offers a solution that directly addresses these daily frustrations while opening new research possibilities.
The hidden costs of traditional synthesis
Research delays due to “unmanufacturable” DNA templates are a common challenge in mRNA therapeutic development, especially when designing constructs with long poly(A) tails.
These tails, made up of repeated adenine bases at the end of mRNA molecules, are essential for enhancing mRNA stability and protein expression. However, traditional chemical DNA synthesis (CDS) is often limited to sequences around 200 nucleotides.
To create the longer DNA templates required for effective mRNA production, researchers must stitch together multiple shorter, chemically synthesized fragments.
However, phosphoramidite chemistry doesn’t join DNA building blocks with perfect efficiency, especially over longer sequences. At each step, some DNA blocks fail to link correctly, which means that producing a 200-nucleotide sequence often results in a mixture where the correctly synthesized, full-length oligonucleotide is in the minority.
As a result, researchers can spend hours troubleshooting failed cloning reactions, repeating experiments due to inconsistent results and dealing with high downstream purification costs, potentially derailing entire therapeutic development programs and compromising the final product’s efficacy.
Modern labs prioritizing sustainability find themselves in a bind as well. The acetonitrile and other harsh solvents required for chemical synthesis generate hazardous waste that’s expensive to dispose of and contradicts green initiatives.
For automated facilities, the infrastructure requirements for safe handling of these chemicals add complexity and cost.
How enzymatic synthesis is transforming research
Unlike traditional phosphoramidite chemistry that relies on harsh chemical reactions, enzymatic DNA synthesis (EDS) uses engineered enzymes to build DNA sequences in mild, aqueous conditions. This biomimetic approach leverages nature’s own machinery – specifically engineered polymerases that can accurately add nucleotides in a controlled, stepwise manner.
The result is a synthesis method that avoids the toxic solvents and harsh conditions of chemical synthesis while maintaining the precision needed for modern molecular biology applications.
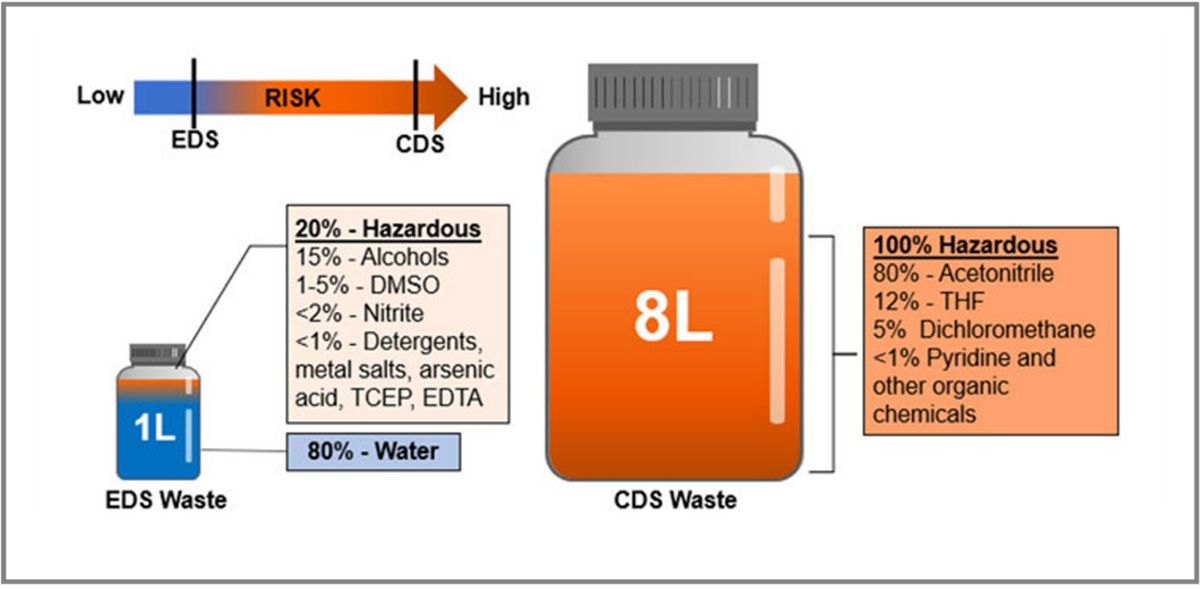
Figure 1: Analysis of waste stream for a benchtop EDS instrument and CDS instrument with the hazardous nature of waste indicated by color (non-hazardous = blue; hazardous = orange). Credit: Simmons et al. 2023.1 CC BY 4.0.
Because EDS operates in conditions that preserve DNA integrity while handling complex sequences, researchers are increasingly reporting successful synthesis outcomes. These include complete in vitro transcription sequences with poly(A) tails up to 400 nucleotides and successful use of templates as long as 500 nucleotides.
Additionally, EDS has made it possible to produce high-GC content fragments that previously required multiple redesign attempts, as well as complex regulatory elements with extensive secondary structures (Table 1).
Table 1: Comparison of enzymatic DNA synthesis and chemical synthesis across four key metrics.
|
|
Enzymatic DNA synthesis |
Chemical synthesis |
|
Turnaround time |
Rapid synthesis of DNA templates in hours to days. |
Limited length and complexity prolong turnaround for full-length templates. |
|
Purity |
Direct synthesis without bacterial amplification or ligation, minimizing errors and unwanted mutations. |
Error rates increase with length, particularly for complex sequences. |
|
Length |
Up to 500 nt and even longer constructs can be assembled using produced oligos. Handles challenging sequences, including long poly(A) tails, ITRs and high GC content. |
Generally limited to ~200 nt, requiring stitching of shorter fragments, which adds complexity and increases error risk. |
|
Sustainability |
Uses aqueous reagents, minimizing hazardous waste and promoting sustainable manufacturing. |
Relies on harsh organic solvents, generating considerable hazardous waste. |
Benchtop EDS systems enable labs to design a construct in the morning and have the DNA ready for cloning by the afternoon, substantially speeding up design-build-test cycles from weeks to days.
The high fidelity of enzymatic synthesis means researchers spend less time screening colonies and more time generating meaningful data. Studies and internal benchmarks show that EDS users experience reduction in screening efforts while maintaining consistent results across experimental replicates and successful first attempts at cloning for >90% of constructs.2
Real value for important applications
The technical capabilities of enzymatic DNA synthesis directly address several persistent challenges in modern molecular biology research (Figure 2).
Figure 2: Key applications of enzymatic DNA synthesis. Credit: DNA Script.
mRNA vaccine and therapeutic engineering
Effective mRNA therapeutics require precise structural elements for stability and translation. The length of the poly(A) tail directly impacts mRNA potency.3 Tails longer than 100 nucleotides significantly enhance stability and expression (Figure 3). Traditional plasmid methods often face challenges due to E. coli recombination with long homopolymeric sequences.
In contrast, EDS can directly synthesize continuous poly(A) tails up to ~400 nucleotides, enabling exploration of novel designs. Grier et al. demonstrated that their linear plasmid vector (pEVL system), used to generate mRNA with extended poly(A) tails, achieved up to a sixfold increase in expression compared to enzymatically polyadenylated mRNA, a particularly important advantage for applications requiring sustained protein production.3
Figure 3: The effect of poly(A) tail length on mRNA potency. Credit: Adapted from Grier et al.3
Adeno-associated viral (AAV) vector development for gene therapy
AAV vectors dominate gene therapy delivery, but production faces persistent bottlenecks. Inverted terminal repeats (ITRs) – palindromic sequences essential for viral packaging – are notoriously difficult to synthesize due to their self-complementary nature. ITR modifications can significantly impact transgene expression.4
EDS reliably produces high-fidelity ITRs in single fragments, eliminating complex workarounds that could compromise vector function, which is critical as the field explores novel serotypes requiring even more complex ITR variants.
Antibody engineering and beyond
EDS streamlines therapeutic antibody development by enabling direct synthesis of full-length single-chain antibody fragments.5 More broadly, EDS handles previously impossible sequences including GC-rich sequences and tandem repeats, removing fundamental barriers in synthetic biology design.
By solving these technical challenges with higher fidelity and faster turnaround, EDS isn’t just improving workflows, it’s enabling more ambitious therapeutic designs and accelerating next-generation treatment development.
Economic impact: Beyond the bench
The advantages of enzymatic synthesis extend beyond technical capabilities, offering significant economic benefits to research and development. By drastically reducing the need for troubleshooting failed experiments, repeated purifications and multiple rounds of construct redesign, EDS directly lowers operational costs associated with reagent waste and researcher time.
The ability to accelerate design-build-test cycles from weeks to days and test 10 times more variants in the same timeframe translates into faster project completion and quicker time-to-market for new therapies. Avoiding hazardous solvent disposal costs associated with traditional methods contributes to more sustainable and economically sound laboratory operations.
While upfront investment in EDS systems may be a consideration, the long-term savings from increased efficiency, reduced material waste and accelerated discovery can lead to a significant return on investment, allowing resources to be reallocated towards more innovative research.
The future is already here
The ability to synthesize previously impossible sequences represents a fundamental shift in research capabilities. As bioengineering pursues increasingly complex designs – multi-domain proteins, precision therapeutics and intricate regulatory elements – traditional synthesis limitations become critical bottlenecks. For researchers pushing therapeutic boundaries, enzymatic synthesis enables entirely new experimental approaches.
Combined with advances in computational design and automation, this technology promises to compress discovery timelines from years to months. The question isn’t whether these capabilities will transform research, but how quickly we can harness them to address urgent therapeutic challenges.
1. Simmons BL, McDonald ND, Robinett NG. Assessment of enzymatically synthesized DNA for gene assembly. Front Bioeng Biotechnol. 2023;11:1208784. doi: 10.3389/fbioe.2023.1208784
2. DNA Script. Gene Assembly. Published August 2022. Accessed November 21, 2025.
3. Grier AE, Burleigh S, Sahni J, et al. pEVL: A linear plasmid for generating mRNA IVT templates with extended encoded poly(A) sequences. Mol Ther Nucleic Acids. 2016;5:e306. doi: 10.1038/mtna.2016.21
4. Shitik EM, Shalik IK, Yudkin DV. AAV-based vector improvements unrelated to capsid protein modification. Front Med. 2023;10:1106085. doi: 10.3389/fmed.2023.1106085
5. Kalantar M, Kalanther I, Kumar S, Buxton EK, Raeeszadeh-Sarmazdeh M. Determining key residues of engineered scFv antibody variants with improved MMP-9 binding using deep sequencing and machine learning. Comput Struct Biotechnol J. 2024;23:3759–3770. doi: 10.1016/j.csbj.2024.10.005
link

