RNA-associated glycoconjugates highlight potential ambiguities in glycoRNA analysis
Mammalian cell culture
HeLa cells were cultured at 37 °C and 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM) (Welgene for Figs. 1b, c, 2b, c, e, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b–f, 4a, 5d,e and Gibco for Supplementary Fig. 2e) supplemented with 10% fetal bovine serum (FBS) (Welgene for Figs. 1b, c, 2b, c, e, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b,c,d,f, 4a and 5d,e and Sigma-Aldrich for Supplementary Fig. 2e) and also supplemented with penicillin–streptomycin (Sigma-Aldrich, Supplementary Fig. 2e). The cells were maintained in 100-mm cell culture dishes (Figs. 1b, c, 2b, c, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b,c,d,f, 4a and 5d,e) with 10 ml of culture media or in T75 flasks (Supplementary Fig. 2e). When reaching confluency, cells were split for subculturing. K562 cells (Supplementary Fig. 5b) were cultured at 37 °C and 5% CO2 in RPMI medium 1640 supplemented with 10% FBS (Cytiva) and penicillin–streptomycin (Sigma-Aldrich) with shaking at 120 rpm.

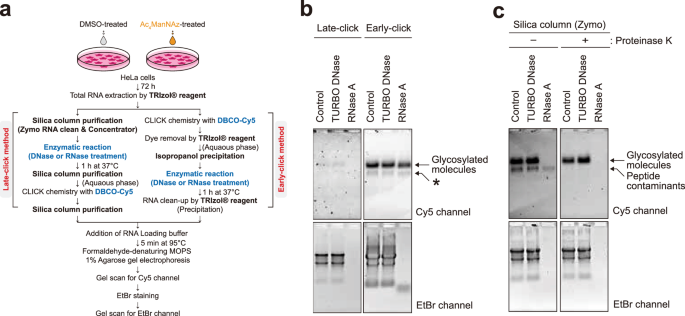
a A schematic comparison of early click and late-click protocols. b Glycan detection by early click and late-click methods. The asterisk indicates the presence of labeled peptide contaminants (see c). c Glycan visualization with or without proteinase K treatment.

a Schematics of the experimental procedure for labeling and visualization. b Glycan detection in Cy5 channel from a denaturing agarose gel without any enzymatic treatment condition. DMSO was used as an Ac4ManNAz untreated control. EtBr channel imaged on the gel scanner shows RNAs were intact. c Glycan visualization in total RNA from HeLa cells or size-fractionated RNAs. d A schematic representation of experiments in Fig. 2e, f. e A comparison of RNA clean-up steps between silica column and TRIzol protocols for glycan visualization. DBCO-Cy5-labeled RNA samples were reacted with TURBO DNase or RNase A or Rapid PNGase F or α2-3,6,8,9 neuraminidase A. The data represent one of three replicates. f Blotting of biotinylated glycosylated molecules treated with various nucleases using streptavidin-conjugated IR800 dyes on the nitrocellulose membrane. Right: gel image of EtBr-stained RNA samples. Left: fluorescent image using IR800-streptavidin in 800-nm channel.
Labeling with MCRs
Stocks of 500 mM N-azidoacetylmannosamine-tetraacylated (Ac4ManNAz) (Sigma-Aldrich, Figs. 1b, c, 2b, c, e, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b,c,d,f,4a and 5b,d,e) were prepared in sterile dimethyl sulfoxide (DMSO) (Sigma-Aldrich). For metabolic labeling, we treated cells with Ac4ManNAz at a final concentration of 100 µM in fresh DMEM supplemented with 10% FBS. After 72 h incubation at 37 °C and 5% CO2, the cells were washed with Dulbecco’s phosphate-buffered saline (PBS) (Welgene, Figs. 1b, c, 2b, c, e, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b,c,d,f, 4a and 5b,d,e) twice and stored in −80 °C until total RNA extraction. For experiments shown in Supplementary Fig. 5b, the conditions for metabolic labeling were the same, except for that penicillin–streptomycin was added in the media. For experiments shown in Supplementary Fig. 2e, of 50 mM stocks of Ac4ManNAz (custom synthesis by Synvenio) were prepared in sterile DMSO (Merck). The metabolic labeling was done with 100 µM Ac4ManNAz in DMEM (Gibco) supplemented with 10% FBS (Sigma-Aldrich) and 1% penicillin–streptomycin. After 48 h at 37 °C and 5% CO2, the cells were washed with PBS (in-house preparation) and then followed by total RNA extraction.
Total RNA extraction with TRIzol reagent
For the experiments shown in Figs. 1b, c, 2b, c, e, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b,c,d,f, 4a and 5d,e, 1 ml of TRIzol reagent (Thermo Fisher Scientific) was added directly onto a washed cell culture dish. The dishes were rocked thoroughly for 10 min at room temperature to lyse and denature all cells. For extracting total RNA from K562 cells (Supplementary Fig. 5b), the cell pellets washed twice with Dulbecco’s PBS were lysed in TRIzol reagent (Thermo Fisher Scientific) with 1 ml for ~107 cells). Homogenized TRIzol-cell mixtures were scraped with cell scraper and then transferred into nuclease-free sterile 1.7 ml microtubes. The tubes were vortexed at least for 5 min until complete homogenization for further denaturation of the intermolecular noncovalent interactions. Phase separation was initiated by adding 200 µl (0.2× volumes) of 100% chloroform (Sigma-Aldrich) into 1-ml TRIzol-dissolved cell mixture and then vortexed thoroughly for complete homogenization. The homogenates were centrifuged at 16,000g for 10 min at 4 °C. The upper (aqueous) phase was carefully removed, transferred into a nuclease-free sterile 1.7 ml and then mixed with equal volume of 100% isopropanol (Fisher Scientific) by vortexing. The mixture was centrifuged at 16,000g for 30 min at 4 °C, and the supernatant was carefully discarded. The pellet was washed with 1 ml of ice-cold 75% ethanol (Sigma-Aldrich) twice and then dried completely. The RNA pellet was dissolved with Milli-Q (Thermo Fisher Scientific, Figs. 1b, c, 2b, c, f, 4b, c, 5b, c and Supplementary Figs. 1, 2b–f, 4a and 5d,e) H2O or Diethyl pyrocarbonate (DEPC)-treated H2O (Thermo Fisher Scientific, Supplementary Fig. 5b), and the concentration was measured using a Nanodrop 2000 ultraviolet–visible light spectrophotometer (Thermo Fisher Scientific).
For experiments shown in Supplementary Fig. 2e, 6 ml of TRI Reagent Solution (Thermo Fisher Scientific) were added directly to the washed T75 cell culture flask. The homogenates were vortexed for 1 min at room temperature followed by incubating the samples 1 min at 37 °C. A total of 0.2× volumes of chloroform were added, and phase separation was performed at 16,000g for 10 min at room temperature. After adding equal volume of isopropanol, mixtures were incubated for 10 min at 4 °C. The RNA was precipitated at 16,000g for 10 min at 4 °C, washed twice with 75% ethanol and dissolved in nuclease-free H2O. To obtain highly pure RNA preparations, the isolated RNA was repurified by adding 1 ml of TRI Reagent Solution (Thermo Fisher Scientific) and repeating the isolation procedure described above.
Copper-free click chemistry and removal of free ligands
Strain-promoted alkyne-azide cycloadditions was performed to probe for azide-containing sialo-conjugates in RNA samples using dibenzocyclooctyne-conjugated cyanine 5 (DBCO-Cy5) (Sigma-Aldrich) dyes or poly-ethylene glycol-spaced dibenzocyclooctyne-biotin (DBCO-biotin, Sigma-Aldrich) as the alkyne-azide cycloaddition. Stocks of 10 mM DBCO-Cy5 or DBCO-biotin were made by dissolving 1 mg of lyophilized DBCO-Cy5 in 82.6 µl or 5 mg of DBCO-biotin in 666.7 µl of DMSO, respectively. A total of 9 µl (typically ~50 µg) of RNA dissolved in H2O were mixed at first with 10 µl of home-made 2× dye-free RNA loading buffer (95% formamide, 25 mM ethylenediaminetetraacetic acid, pH 8.0) and added with 1 µL of 10 mM (for final 500 µM) DBCO-Cy5 or DBCO-biotin in a microtube. The samples were incubated at 55 °C for 10 min. The reaction was stopped by adding 1 ml of TRIzol reagent (Thermo Fisher Scientific) and 200 µl of chloroform (Sigma-Aldrich). Alternatively, for experiments performed in Supplementary Fig. 2e, dye removal was achieved by adding 80 µl of DEPC-treated H2O, 300 µl of TRIzol LS Reagent (Thermo Fisher Scientific) and 80 µl of chloroform (Merck). The samples were centrifuged at 16,000g for 10 min at 4 °C or, for experiments shown in Supplementary Fig. 2e, at 16,000g for 5 min at room temperature. The upper (aqueous) phase was carefully removed and transferred into a nuclease-free sterile tube. The samples were mixed with equal volume of 100% isopropanol by vortexing, subsequently centrifuged at 16,000g for 30 min at 4 °C, and the supernatant was carefully discarded. The pellet was washed with 1 ml of ice-cold 75% ethanol twice and then dried completely. The RNA pellet was dissolved with Milli-Q H2O (Figs. 1b, c, 2b, c, e, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b–f, 4a and 5d,e) or DEPC-treated H2O (Supplementary Fig. 5b), and the concentration was measured using the ultraviolet–visible light spectrophotometer.
Enzymatic reactions
Typically, enzymatic reactions were performed with 10 µg (Figs. 1b, c, 2b, c, f, 4b, c and 5b, c and Supplementary Figs. 1, 2b,c,d,f, 4a and 5b,d,e) or 12 µg (Supplementary Fig. 2e) of labeled RNA at 37 °C. To digest RNA, samples were treated with 0.5 µl of RNase A (DNase and protease-free, 10 mg/ml) (Thermo Fisher Scientific, Figs. 1b, c, 2c, e, f, 4b, c and 5b, c and Supplementary Fig. 1, 2f, 4a and 5b,d), PureLink RNase A (20 mg/ml) (Thermo Fisher Scientific, Supplementary Fig. 5b) or 1 µL of RNase T1 (Thermo Fisher Scientific, Fig. 2f and Supplementary Fig. 2f) or 1 µl of RNase cocktail (500 U of RNase A and 20,000 U of RNase T1 per milliliter) (Thermo Fisher Scientific, Fig. 2f and Supplementary Fig. 2f) prepared in 1.5 µl of 10× TURBO DNase buffer (Thermo Fisher Scientific adjusting the final volume with Milli-Q H2O to 15 µl). To degrade DNA, samples were treated with 0.5 µl of TURBO DNase (2000 U/ml) (Thermo Fisher Scientific, Figs. 1b, c and 2c, e, f and Supplementary Figs. 1 and 2c,d,f) or DNase I (2,000 U/ml) (Thermo Fisher Scientific, Supplementary Fig. 2e) prepared in 1.5 µL of 10× DNase buffer adjusting the final volume with Milli-Q H2O to 15 µl. To digest both RNA and DNA, the samples were treated with 1 µl of benzonase (250,000 U/ml) (Merck, Fig. 2f and Supplementary Fig. 2f) or 1 µl of nuclease P1 (100,000 U/ml) (New England Biolabs, Fig. 2f and Supplementary Fig. 2f) prepared in 1.5 µl of 10× TURBO DNase buffer or nuclease P1 buffer adjusting the final volume with Milli-Q H2O to 15 µl. To digest RNA in DNA/RNA hybrids, samples were treated with 1 µl of RNase H (5000 U/ml) (New England Biolabs, Fig. 2f and Supplementary Fig. 2f) prepared in 1.5 µl of 10× RNase H buffer adjusting the final volume with Milli-Q H2O to 15 µl. To digest N-glycans, 0.5 µl of Rapid PNGase F (New England Biolabs, Fig. 2f and Supplementary Fig. 2f) were added with 1.5 µl of 10× PNGase F buffer (New England Biolabs) in the sample by adjusting with Milli-Q H2O to total 15 µl. To cut sialic acid moieties, 0.5 µl of α2-3,6,8,9 neuraminidase A (New England Biolabs, Fig. 2f and Supplementary Fig. 2f) were added with 1.5 µl of 10× GlycoBuffer 1 (New England Biolabs) in the sample by adjusting with Milli-Q H2O to total 15 µl. To digest proteins, 1 µl of proteinase K (Roche, 20 mg/ml dissolved in Milli-Q H2O, Figs. 1c, 2c, e, f, 4b, c and 5b, c and Supplementary Fig. 1, 2c,d,f, 4a and 5d, e; Thermo Fischer Scientific, 20 mg ml, Supplementary Fig. 2e) was added either with 1.5 µl of 10× TURBO DNase buffer in the RNA sample by adjusting with Milli-Q H2O to total 15 µl or directly in the precedent enzymatic reactant. The incubation was done for 30 or 60 min in cases, but all the results always exhibited complete protein digestion.
RNA fragmentation
DBCO-Cy5-labeled RNA was fragmented using Ambion 10X RNA Fragmentation Reagent (Thermo Fisher Scientific). The samples were incubated in 1× RNA Fragmentation Reagent at 75 °C for 15 min for mild reaction (Supplementary Fig. 5d) and at 95 °C for 2 h for complete fragmentation (Supplementary Fig. 5e). Fragmented RNA samples were immediately mixed with 2× volumes of RNA binding buffer (RBB; RNA Clean and Concentrator-5 kit, Zymo Research) and various volumes of 100% ethanol for each sample for the final 40%, 50%, 60% and 70% ethanol, as indicated in Supplementary Fig. 5d,e.
RNA clean-up by acidic phenol-chloroform extraction
For experiments in Figs. 1b and 2e, f and Supplementary Fig. 2d,e,f, enzymatically digested samples were mixed thoroughly with 1 ml of TRIzol reagent and 200 µl of 100% chloroform (Sigma-Aldrich) for 10 min at room temperature. The homogenates were centrifuged at 16,000g for 10 min at 4 °C. The upper phase was carefully removed, transferred into a nuclease-free sterile 1.7-ml tube and then mixed with 1 µl of linear acrylamide (Thermo Fisher Scientific) as a coprecipitant and equal volume of 100% isopropanol by vortexing, subsequently centrifuged at 16,000g for 30 min at 4 °C, and the supernatant was carefully discarded. The pellet was washed with 1 ml of ice-cold 75% ethanol twice and then dried completely. The pellet was dissolved with Milli-Q H2O. For experiments in Supplementary Fig. 2e, 20 µg linear acrylamide and TRI Reagent Solution (Thermo Fisher Scientific) were used.
RNA clean-up and size fractionation by silica-based column purification
A total of 16 µl of proteinase K-digested samples were mixed with 34 µl of Milli-Q H2O. A total of 100 µl of RBB and 150 µl of 100% ethanol (final 50% ethanol) were added by reverse pipetting and vortexed thoroughly. For experiments shown in Supplementary Fig. 2e, the final percentage of EtOH was 60%. The mixtures were transferred into the Zymo-SpinTM IC column in a 2 ml collection tube. The columns were centrifuged at 16,000g for 30 s at room temperature, and the flow-through was discarded. A total of 400 µl of RNA prep buffer (RPB; RNA Clean and Concentrator-5 kit, Zymo Research) were added into the column and then centrifuged at 16,000g for 30 s at room temperature followed by discarding the flow-through. A total of 700 µl of RNA wash buffer (RWB; RNA Clean and Concentrator-5 kit, Zymo Research) were added into the column and then centrifuged at 16,000g for 30 s at room temperature, followed by discarding the flow-through. A total of 400 µl of RWB were added into the column and then centrifuged at 16,000g for 30 s at room temperature, followed by discarding the flow-through. To ensure complete removal of the RWB, samples were again centrifuged at 16,000g for 1 min at room temperature. The columns were carefully transferred into a new nuclease-free sterile 1.7-ml tube, and 15 µl of Milli-Q H2O or DEPC-treated H2O was directly applied to the column matrix and incubated for 3 min. The elution was done by centrifugation at 16,000g for 3 min at room temperature.
For size fractionation of small (smaller than about 200 nucleotides) versus large (larger than about 200 nucleotides) RNAs, samples were mixed with equal volume of 50% RBB in 100% ethanol. The mixture was applied onto the Zymo-SpinTM IC Column and centrifuged at 16,000g for 1 min at room temperature. Large RNAs bound in the column were purified as described above. Small RNAs in the flow-through were mixed with equal volume of 100% ethanol and then purified as described above.
Photochemical labeling of fragmented RNA with amine-DBCO and click chemistry
The purified RNA sample was fragmented using the NEBNext Magnesium RNA Fragmentation Module (E6150S) following protocols recommended by the vendor and purified using Zymo RCC-25. Amine-DBCO (CAS no. 1255942-06-3) methylene blue (CAS no. 61-73-4) was added to a solution of 50 µg RNA at a final concentration of 1 mM and 100 µM, respectively. The solution was exposed to red light (50 W light-emitting diode chip on board) on ice for 15 min with lid open. The resulted reaction mixture was then purified using TRIzol in combination with Zymo RCC-25 following the vendor-provided protocol. The purified DBCO-functionalized RNA (10 µg per reaction) was then clicked with either Cy5-azide (Jena Bioscience, CLK-1177) or the AlexaFluor-555-labeled sialoglycan-azide at 200 and 60 µM, respectively, at 50 °C for 10 min. The clicked RNA product was then purified with TRIzol in combination with Zymo RCC-10 following the vendor’s protocol. The RNA samples were resolved with 1% denaturing agarose gel and fluorescently imaged under a GelDoc scanner, after which the gel was stained with ethidium bromide (EtBr) and scanned again for total RNA.
To make AlexaFluor-555-labeled sialoglycan-azide, G2 glycoform N-glycan linked with an azido-asparagine (final concentration 120 µM) was mixed with 10 µg ST6Gal1 and 200 µM cytidine monophosphate–sialic acid prefunctionalized with AlexaFluor-555. The sialylation reaction was incubated at 37 °C for 48 h and was used for the click chemistry with RNA directly without further purification.
Denaturing gel electrophoresis, membrane transfer and blotting
Typically, formaldehyde-denaturing 1% agarose gels were prepared by thoroughly mixing 0.5 g of agarose powder (Lonza) with 45 ml of Milli-Q H2O in an Erlenmeyer flask. The agarose was melted by heating in the microwave oven and then cooled to 55–60 °C. A total of 5 ml of 10× NorthernMax denaturing gel buffer (Thermo Fisher Scientific) were added and mixed by swirling in the fume hood. The gel mixture was casted following the instruction provided by the casting apparatus. To prepare loading samples, the samples were mixed with equal volume of 2× dye-free RNA loading buffer and then incubated at 95 °C for 10 min. The gel was resolved in 1× NorthernMax running buffer (Thermo Fisher Scientific) at 100 V for 40–50 min. For visualization of the Cy5 fluorescence, the gel was rinsed briefly with Milli-Q H2O and scanned using the gel imaging system (Bio-Rad ChemiDoc XRS+) in the Cy5 filter channel. For EtBr scanning, the Cy5 scanned gel was stained in the water-dissolved UltraPure EtBr (Thermo Fisher Scientific) solution for 30 min at room temperature by rocking. The gel was rinsed with Milli-Q H2O for 30 min at room temperature by rocking and then scanned in the gel imaging system.
For experiments shown in Supplementary Fig. 2e, 1 g of agarose powder (Roche) was dissolved in 72 ml of Milli-Q H2O. A total of 10 ml of 10× 3-(N-morpholino)propanesulfonic acid (MOPS) buffer (200 mM MOPS, 50 mM sodium acetate·3H2O, 10 mM ethylenediaminetetraacetic acid, pH 7.0) and 18 ml 37% formaldehyde (Merck) were added and mixed. Purified, enzyme treated, RNA samples were incubated at 95 °C for 5 min, followed by a quick transfer and 5 min incubation on ice before gel electrophoresis at 90 V. Cy5 fluorescence was visualized using the Amersham Typhoon scanner (GE Healthcare).
For membrane transfer, the electrophoresed gel was scanned in the Cy5 channel and then immediately subjected to the transfer instead of EtBr staining, as the EtBr emission was strongly overlapped with Cy5 visualization during the downstream membrane scanning. NorthernMax transfer buffer (Thermo Fisher Scientific) was used following the manufacturer’s instruction for 2 h. For nylon membranes, Zeta-Probe GT (Bio-Rad) or BrightStar-Plus (Thermo Fisher Scientific) membranes were used. For nitrocellulose membranes, Hybond-C (Cytiva) or Amersham Protran (Sigma-Aldrich) membranes were used. The transferred membranes were rinsed briefly with Milli-Q H2O and scanned immediately in the Cy5 channel using Bio-Rad ChemiDoc XRS+.
After transfer of the gel run with biotinylated samples, the membranes were subjected to blocked with Odyssey blocking buffer, PBS (Li-Cor Biosciences) for 45 min at room temperature, by skipping EtBr staining and fluorescent imaging. After blocking, the membranes were stained for 30 min at room temperature with streptavidin-conjugated IR800 (Li-Cor Biosciences), which was diluted to 1:5,000 in Odyssey blocking buffer. Excess IR800-streptavidin was removed from the membranes by four washes with 0.1% Tween-20 (Sigma-Aldrich) in 1× PBS for 10 min each at room temperature. The membranes were finally washed once with 1× PBS to remove residual Tween-20 before scanning. Fluorescent signals from membranes were scanned on Odyssey Li-Cor Sa scanner (Li-Cor Biosciences) with the software set to autodetect the signal intensity for both 700 and 800 channels. After scanning, the images were adjusted to appropriate contrasts with the Li-Cor software (when appropriate) in the 800 channel and exported.
link





.jpg)